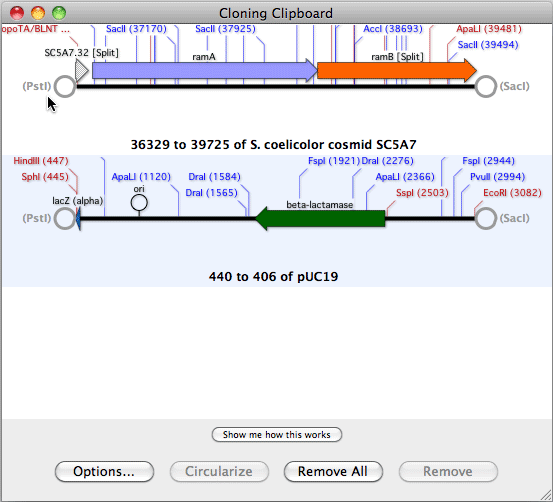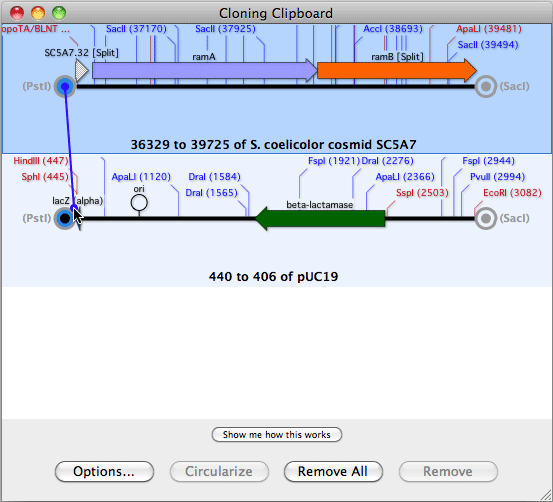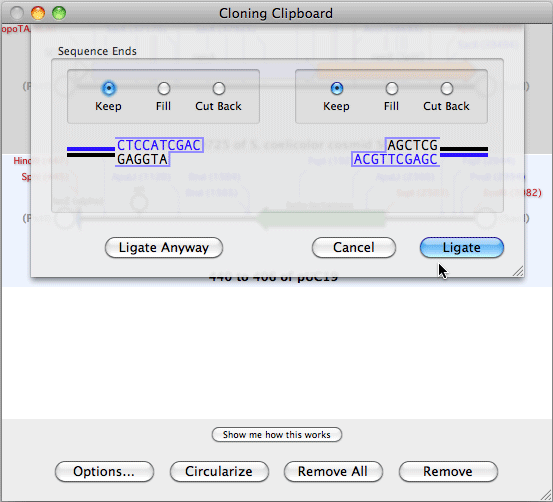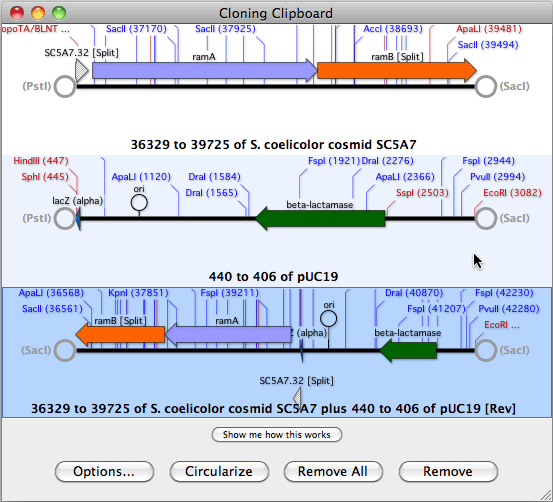 
Initially, the sequence residues are displayed in italics to indicate they have not been aligned.Ĭopyright © 2022 MacVector, Inc. The imported sequences are displayed below the reference sequence. You can then add one or more sequences to the window by clicking on the "+" button. 
This is displayed along the top of the window. The key to this functionality lies in the main Align to Reference editor. The trial version of MacVector includes all of the sample files you will need to follow the tutorial. There is a detailed Sequence Confirmation Tutorial that provides far more information on this functionality that can be downloaded here. A typical example would be in resequencing, e.g. The second use is cDNA Alignments, which allows you to align mRNA, cDNA or EST sequences against a genomic template. Align to Reference Use this tool if you have a reference sequence and you want to align one or more DNA sequences against it. There are two main uses for this: Sequence Confirmation is similar to sequence assembly, except that it requires the use of a known reference sequence as a scaffold. MacVector has a unique Align to Reference interface that lets you align one or more files against a reference sequence. A common request, especially in our recent survey, is to align existing primer sequences against a template sequence.Sequence Analysis Tools for Molecular Biologists There are many ways to do this in MacVector, depending on what your requirements are Using the Find dialogįor quickly finding a single primer in a sequence the Find dialog is the first point of call. This allows you to find any sequence, whether it binds to the complementary strand, it is reversed or both. It also allows you to state which end of the sequence to start from and in which direction to scan the sequence (not necessarily obvious!). The Find dialog (see below) is a little daunting to look at first (in fact due to recent user feedback we will hiding most of the functionality in the next release). (i) open up your sequence and use the menu option EDIT > FIND or use the key combination CMD – F However, it is very powerful and in most cases you do not need to change anything as the default settings will work for the majority of cases. When to use: When you need to find a single primer very quickly and do not need to store the results (ii) Enter your primer sequence in the FIND box and click FIND. (iv) Change the drop down menu to Sequence Confirmation then change these parameters: (i) Open the template file and go to ANALYZE > ALIGN TO REFERENCE If you want to quickly align a large set of primers against a template sequence, then as long as each primer is in a separate MacVector file, or a multi sequence fasta file then you can use Sequence Confirmation in the Align to Reference function: Limitations: You can only find a single primer. – If you suspect your primers may not be a perfect match then reduce SCORE THRESHOLD until your primer aligns. The resulting alignment will show your primers aligned against the template. Select the sequence entitled SequenceSampleand click on the Open button. You can switch to the Map view to show a graphical overview of all your primers and where they are located on the sequence. The first step in the tutorial is to open and explore the sample sequence to be used in the analysis A standard MacVector nucleic acid sequence window will open Select File Openand navigate to the /Tutorial Files/Align To Reference/Sequence Confirmation/folder. When to use: When you need to find many primers over a large sequenceīenefits: Easy to visualize a great quantity of primers against a template You can use the Editor view for a sequence level representation of the primer aligned against the template. Limitations: Your primer sequences need to be in a file already. This function is fairly easy to use and gives a large amount of detail about your primers. (i) Open your sequence and go to PRIMERS > Test PCR Primer Pairs For example secondary structure, what size the product will be and even the most ideal Tm for your PCR run. (ii) Copy and paste your two sequences in the two boxes. Note you will need to have pasted them into an external application. MacVector makes it simple to import or create sequences, automatically annotate sequences, design primers and store them in a database, subclone into cloning vectors, align, assemble reads, nd SNPs, simulate agarose gels, run protein analyses and a wide variety. (iii) Click APPLY and see your primers detailed statistics. MacVector, the leading sequence analysis package for the Mac, takes full advantage of the Mac’s easy to use interface. Primer 2: score 20, mismatches 0, lower strand 1592 to 1573 The 3' end of the primer binds within the product primer 1: score 20, mismatches 0, upper strand 1055 to 1074 Major product size: 538 bp Product details: (iv) Click OK to see the full statistics on the primers and product.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
